MPICWang2013
To discover sequence differences responsible for adaptation to diverse environments across the native range of Arabidopsis thaliana, the Max Planck Institute for Developmental Biology (MPI) and Monsanto Company sequenced 343 wild inbred accessions from 22 countries, with 9- to 112-fold genome coverage using Illumina's sequencing-by-synthesis (SBS) technology.
Paired-end short reads (100 bp) were analyzed using our software pipeline SHORE to identify the SNPS and 1-3bp deletions.
Regional distribution
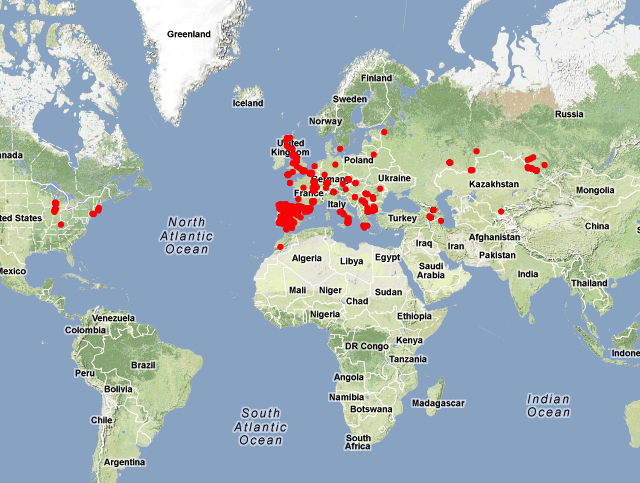
Accessions
We collected strains from 22 countries:
| Number of Strains | Region |
|---|---|
| 87 | ESP |
| 46 | GER |
| 35 | FRA |
| 29 | UK |
| 31 | RUS |
| 21 | USA |
| 23 | ITA |
| 17 | CZE |
| 14 | BUL |
| 6 | POR |
| 6 | ROU |
| 3 | GEO |
| 6 | ARM |
| 4 | SRB |
| 3 | SVK |
| 3 | GRC |
| 2 | CRO |
| 2 | AUT |
| 1 | UZB |
| 1 | SWE |
| 1 | SUI |
| 1 | MAR |
| 1 | #N/A |
For more information please see here.
Data Center
Browse all data in our Data Center and see the Readme for more information on released data files and formats.
Contact
For any questions concerning the data contact Congmao Wang, for any problems downloading/accessing the data contact Joffrey Fitz.